Sharing innovative technologies and research protocols
Researchers with the University of Washington Superfund Research Program work to share their innovative technologies and protocols with the world. The approaches they develop are aimed at detecting and remediating harmful contaminants in the environment and better understanding the effects of these contaminants in the body and in ecological systems. By sharing these approaches widely, we hope to help improve human and environmental health for all.
Fish genes
The Gallagher lab (Project 1) has collaborated with the John Kauwe lab at Brigham Young University to characterize pollutant detoxification genes in the bonefish. This project extends the Gallagher lab Superfund studies targeting salmon and zebrafish to an important tropical fish threatened by anthropogenic activities and potential climate change.
Highly sensitive portable monitoring device
Clement Furlong, a Principal Investigator on Project 2, has helped to develop a highly sensitive, portable monitoring device. The device can be used to provide near real-time measurements of almost any substance for which an immunoassay can be developed. Furlong and his collaborators have partnered with the National Oceanic and Atmospheric Administration and the Monterey Bay Aquarium Research Institute to measure algal toxins. Furlong has also partnered with the U.S. Food and Drug Administration to measure shellfish toxins.
In 2020, Furlong published a paper with colleagues demonstrating the ability of a monitoring device that his lab helped develop to provide rapid, simple, accurate, low-cost, point-of contact diagnosis of tuberculosis (TB) (Kahng et al. 2020). TB is one of the most serious diseases worldwide and accurate, low costs, rapid diagnosis has proven elusive in the past. So this application of the device meets an important need. The TB research was conducted with support from the U.S. Army. Other researchers on the paper were from the UW Department of Mechanical Engineering, the UW School of Engineering and Computer Science, the UW Departments of Medicine-Division of Medical Genetics, and UW Genome Sciences.
Also in 2020, Furlong and colleagues received a grant from the National Institutes of Health (NIH) through its Rapid Acceleration of Diagnostics Initiative (RADx) to prototype a similar protocol for rapid screening of COVID-19 from nasal swab samples. The monitoring device is expected to have 95% sensitivity, 95% specificity and take 15 minutes to provide results, at a cost of 10 dollars per individual sampled. The investigators plan to have the sensor ready for use within one year. They will develop a business plan with the help of UW CoMotion.
Sampling for arsenic in lakes
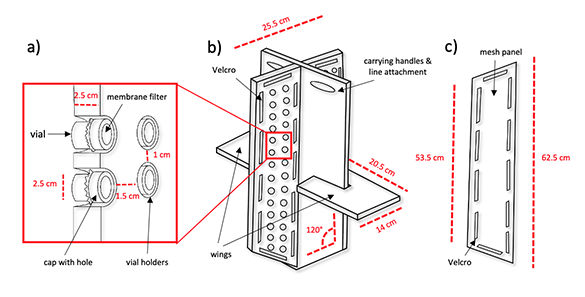
James Gawel and other researchers on Project 4 shared technology that they developed to sample dissolved arsenic with colleagues at the University of Puget Sound. The sampling devices they developed are called "porewater peepers" and are designed to measure small scale (4cm) differences in dissolved arsenic concentration (or other element) above and below the water-sediment interface. This data is critical for estimating the transport of contaminants from sediments to overlying waters.
The porewater peepers were built by UW SRP researchers with the help of students at Bellarmine High School in Tacoma, WA. Along with the loan of equipment, UW SRP researchers provided a demonstration of how to set up and delay the instruments. Jeff Tepper of the University of Puget Sound began deploying the porewater peepers in local lakes starting in July of 2020.
Gawel and trainee Ken Burkhart also developed a low-cost, effective device to sample periphyton from lakes contaminated with arsenic. The UW SRP has been contacted by Summer Gonsalves of the Narragansett Indian Tribe who is interested in employing the technology to look at fish and shellfish contamination in the tribal waters of Narragansett Bay.

